Research should be impactful, novel and exciting. What will you discover?
Precision cancer immunotherapy
A set of new cancer therapies rely on the targeting of markers present at the surface of cancer cells using chimeric antigen receptor T cells, antibody-drug conjugates or vaccines. The application of these revolutionary approaches is limited in part by the lack of cancer-specific markers to target.
Aberrant RNA splicing is a hallmark of all cancer types (e.g. Jha, Quesnel-Vallières et al., 2022). We developped computational tools to identify cancer-specific antigens originating from aberrant splicing variations.
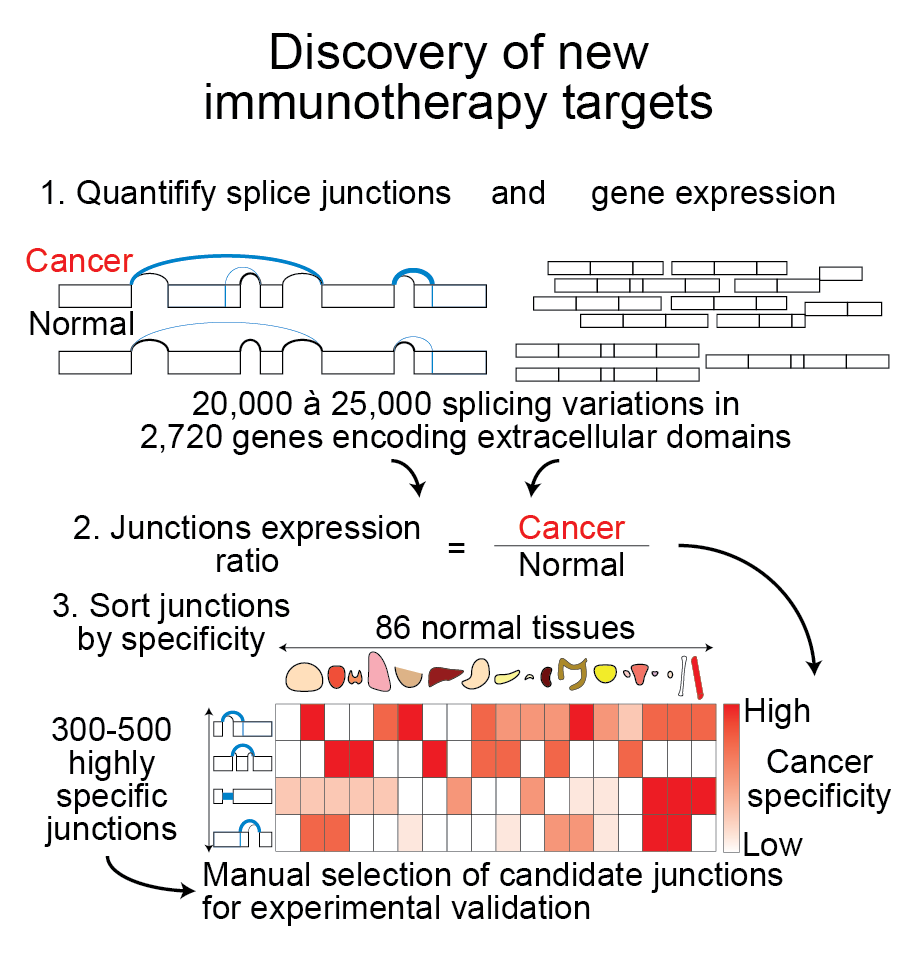 We currently focus our efforts on the experimental validation of
candidates that appear to be prime therapeutic targets
in high-risk cancers such as pancreatic cancer, cholangiocarcinoma,
acute myeloid leukemia and rare forms of pediatric sarcoma and carcinoma.
We currently focus our efforts on the experimental validation of
candidates that appear to be prime therapeutic targets
in high-risk cancers such as pancreatic cancer, cholangiocarcinoma,
acute myeloid leukemia and rare forms of pediatric sarcoma and carcinoma.
Blood-specific RNA splicing regulation
Sets of alternative RNA splicing events are co-regulated to shape the development and function of the nervous system. We previously studied such programs and their regulators (e.g. Quesnel-Vallières et al., 2015). In addition to explaining important molecular mechanisms underlying the complexity of the brain, these alternative splicing programs can be involved in neuronal disorders and diseases (e.g. Quesnel-Vallières et al., 2016).
Similar to components of the nervous system, immune cells display remarkable cellular, molecular and environmental diversity. We believe that a part of this complexity is enabled by hematopoietic-specific alternative splicing events. Our computational analyses have highlighted a number of putative splicing regulators that are specifically expressed in subsets of immune cell types.
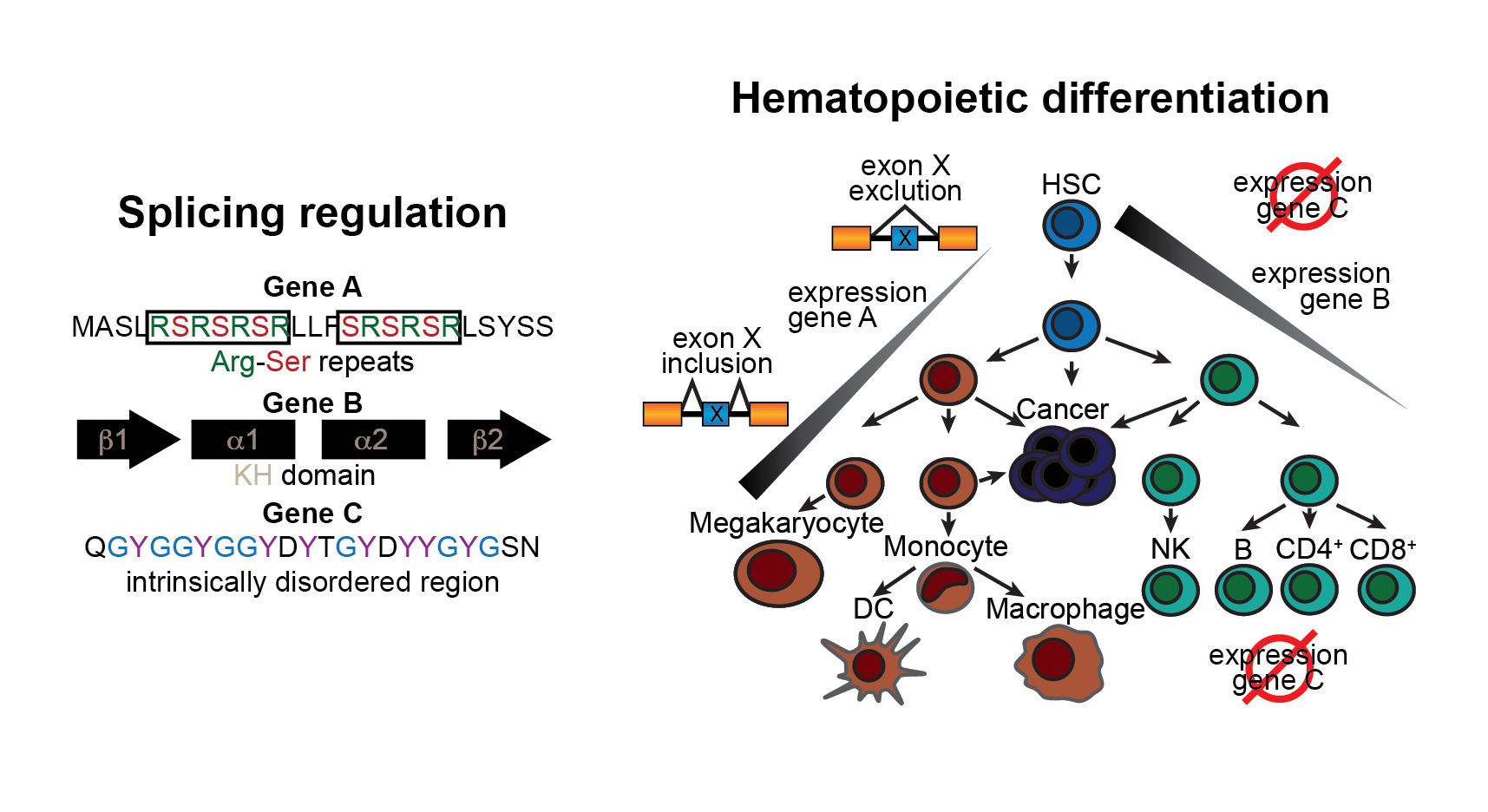
We now aim to confirm the role of top candidates in the regulation of splicing in immune cells and characterize the function of dedicated hematopoietic splicing programs.
What is a gene? What are the evolutionary drivers of gene regulation?
Sometimes we need to ask big questions. You might have some of those questions yourself. Let’s chat.